In a previous post, I described linear discriminant analysis (LDA) using the following Bayes’ rule
where is the prior probability of membership in class
and the distribution of the predictors for a given class
,
, is a multivariate Gaussian,
, with class-specific mean
and common covariance matrix
. We then assign a new sample
to the class
with the highest posterior probability
. This approach minimizes the total probability of misclassification [1].
Fisher [2] had a different approach. He proposed to find a linear combination of the predictors such that the between-class covariance is maximized relative to the within-class covariance. That is [3], he wanted a linear combination of the predictors that gave maximum separation between the centers of the data while at the same time minimizing the variation within each group of data.
Variance decomposition
Denote by the
matrix with
observations of the
-dimensional predictor. The covariance matrix
of
can be decomposed into the within-class covariance
and the between-class covariance
, so that
where
and is the
matrix of class indicator variables so that
if and only if case
is assigned to class
,
is the
matrix of class means,
is the
matrix where each row is formed by
and
is the total number of classes.
Fisher’s approach
Using Fisher’s approach, we want to find the linear combination such that the between-class covariance is maximized relative to the within-class covariance. The between-class covariance of
is
and the within-class covariance is
.
So, Fisher’s approach amounts to maximizing the Rayleigh quotient,
[4] propose to sphere the data (see page 305 of [4] for information about sphering) so that the variables have the identity as their within-class covariance matrix. That way, the problem becomes to maximize subject to
. As we saw in the post about PCA, this is solved by taking
to be the eigenvector of B corresponding to the largest eigenvalue.
is unique up to a change of sign, unless there are multiple eigenvalues.
Dimensionality reduction
Similar to principal components, we can take further linear components corresponding to the next largest eigenvalues of . There will be at most
positive eigenvalues. The corresponding transformed variables are called the linear discriminants.
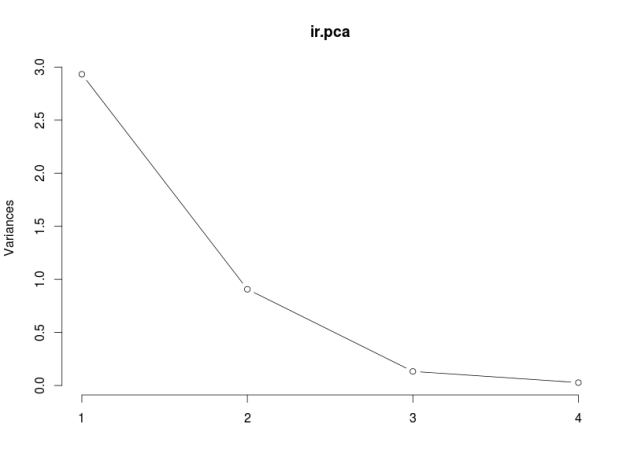
Note that the eigenvalues are the proportions of the between-class variance explained by the linear combinations. So we can use this information to decide how many linear discriminants to use, just like we do with PCA. In classification problems, we can also decide how many linear discriminants to use by cross-validation.
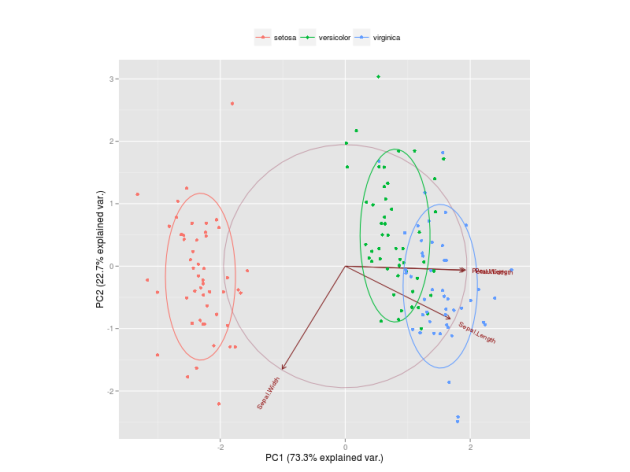
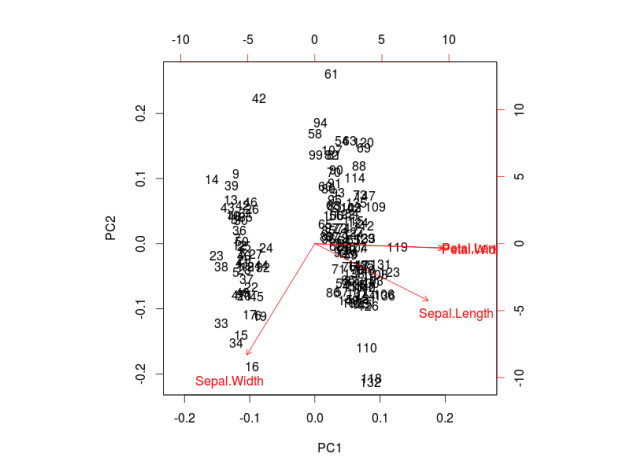
Reduced-rank discriminant analysis can be useful to better visualize data. It might be that most of the between-class variance is explained by just a few linear discriminants, allowing us to have a meaningful plot in lower dimension. For example, in a three-class classification problem with many predictors, all the between-class variance of the predictors is captured by only two linear discriminants, which is much easier to visualize than the original high-dimensional space of the predictors.
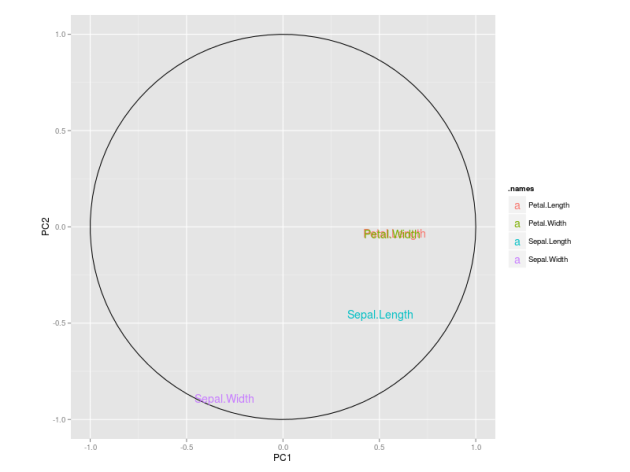
Regarding interpretation, the magnitude of the coefficients can be used to understand the contribution of each predictor to the classification of samples and provide some understanding and interpretation about the underlying problem.
RRDA and PCA
It is important to emphasize the difference between PCA and Reduced-rank DA. The first computes linear combinations of the predictors that retain most of the variability in the data. It does not make use of the class information and is therefore an unsupervised technique. The latter computes linear combinations of the predictors that retain most of the between-class variance in the data. It uses the class information of the samples and therefore belongs to the group of supervised learning techniques.
References:
[1] Welch, B. L. (1939). Note on discriminant functions. Biometrika, 31(1/2), 218-220.
[2] Fisher, R. A. (1936). The use of multiple measurements in taxonomic problems. Annals of eugenics, 7(2), 179-188.
[3] Kuhn, M. and Johnson, K. (2013). Applied Predictive Modeling. Springer.
[4] Venables, W. N. and Ripley, B. D. (2002). Modern applied statistics with S. Springer.